Global atlas of soil microbiomes
At the Helmholtz Centre for Environmental Research, scientists have compiled 15,000 metagenome data sets from soil samples of various origins according to uniform standards.

The soil is teeming with microorganisms. Researching them is not easy because many of them cannot yet be cultivated in the laboratory. Therefore, researchers often record so-called metagenomes - the entirety of the genes of the microorganisms in a sample. More than 200,000 such metagenomes are available in public databases. But there is a problem: the data sets in them are not subject to a uniform standard, making it difficult to put the data to use. A team from the Helmholtz Centre for Environmental Research (UFZ) has now processed 15,000 of these data records according to uniform standards and merged them into a new database. This was published in the scientific journal "Nucleic Acids Research".
Standardized metadata
The data are taken from the databases "MG Rast" and "Sequence Read Archive". However, the datasets are often incomplete and not uniformly marked. For example, information on the temperature of the soil sample can be stored in Kelvin, Fahrenheit or Celsius. "This makes it more difficult for interested users to further process the data," says Ulisses Nunes da Rocha, microbial ecologist at the UFZ. For the new "TerrestrialMetagenomeDB" database, the researchers have therefore standardized all metadata such as temperature, pH value and geographical coordinates according to an existing standardization methodology.
The "TerrestrialMetagenomeDB" metagenome database can also break down its data sets geographically.
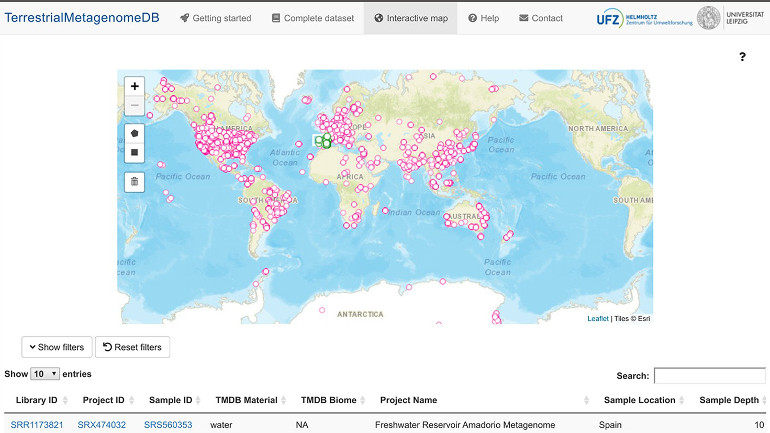
The prepared data sets come from 84 countries and include samples from forest, grassland and rocky soils. Users of the database can use various filters to identify relevant metagenomes. It is also possible to filter by geographical criteria using an interactive world map to which the data sets are assigned. Scientists can thus use the database to check whether certain experiments - for example on CO2 fixation or the influence of pesticides - have been carried out previously and whether comparative data are available. The UFZ researchers themselves want to use the database to carry out analyses of soil microbiomes on a global scale.
Automatic updates
The database, which was launched in November 2019, will continue to grow in the future: New entries in "MG Rast" and "Sequence Read Archive" will be automatically transferred to the new database twice a year, provided they meet the standards.
bl/um